Ascomycota
Sac Fungi
John W. Taylor, Joey Spatafora, and Mary Berbee

This tree diagram shows the relationships between several groups of organisms.
The root of the current tree connects the organisms featured in this tree to their containing group and the rest of the Tree of Life. The basal branching point in the tree represents the ancestor of the other groups in the tree. This ancestor diversified over time into several descendent subgroups, which are represented as internal nodes and terminal taxa to the right.

You can click on the root to travel down the Tree of Life all the way to the root of all Life, and you can click on the names of descendent subgroups to travel up the Tree of Life all the way to individual species.
For more information on ToL tree formatting, please see Interpreting the Tree or Classification. To learn more about phylogenetic trees, please visit our Phylogenetic Biology pages.
close boxIntroduction
The Ascomycota, or sac fungi, is monophyletic and accounts for approximately 75% of all described fungi. It includes most of the fungi that combine with algae to form lichens, and the majority of fungi that lack morphological evidence of sexual reproduction. Among the Ascomycota are some famous fungi: Saccharomyces cerevisiae, the yeast of commerce and foundation of the baking and brewing industries (not to mention molecular developmental biology), Penicillium chrysogenum, producer of penicillin, Morchella esculentum, the edible morel, and Neurospora crassa, the "one-gene-one-enzyme" organism. There are also some infamous Ascomycota, a few of the worst being: Aspergillus flavus, producer of aflatoxin, the fungal contaminant of nuts and stored grain that is both a toxin and the most potent known natural carcinogen, Candida albicans, cause of thrush, diaper rash and vaginitis, and Cryphonectria parasitica, responsible for the demise of 4 billion chestnut trees in the eastern USA (Alexopoulos et al., 1996). Asexual Ascomycota, such as Penicillium or Candida species, used to be classified separately in the Deuteromycota because sexual characters were necessary for Ascomycota classification. However, the comparison of nucleic acid sequence, as well as nonsexual phenotypic characters, have permitted the integration of asexual fungi into the Ascomycota (Taylor, 1995). The Deuteromycota is no longer recognized as a formal taxon in fungal systematics.
Characteristics
The shared derived character that defines the Ascomycota is the ascus. It is within the ascus that nuclear fusion and meiosis take place. In the ascus, one round of mitosis typically follows meiosis to leave eight nuclei, and eventually eight ascospores. Ascospores are formed within the ascus by an enveloping membrane system, which packages each nucleus with its adjacent cytoplasm and provides the site for ascospore wall formation. These membranes apparently are derived from the ascus plasma membrane in the Pezizomycotina and the nuclear membrane in the Saccharomycotina (Wu and Kimbrough, 1992; Raju, 1992).
In hyphal Ascomycota (left), the youngest, terminal hyphal segments develop into 8-spored asci. In yeasts (right) a single cell simply becomes the ascus, often with just 4 spores. Asci of a hyphal Ascomycota (Pezizomycotina), Podospora , © R. Vilgalys 1996. Ascus of a yeast (Saccharomycotina), Saccharomyces, © J. Taylor 1996.
At the time they are released from the ascus, the thick-walled haploid ascospores are resistant to adverse environments. But, given the right conditions, they will germinate to form a new haploid fungus.
The body of Ascomycota is shared by other fungi and consists of a typical eukaryotic cell surrounded by a wall. The body can be a single cell, as in yeasts, or a long tubular filament divided into cellular segments, which is called a hypha (plural, hyphae). Both yeasts and hyphae have cell walls made of varying proportions of chitin and beta glucans (Wessels, 1994).
Nutrition and Symbioses
Like other fungi, Ascomycota are heterotrophs and obtain nutrients from dead or living organisms (Griffin, 1994; Carroll and Wicklow, 1992). If water is present, as saprotrophs they can consume almost any carbonaceous substrate, including jet fuel (Amorphotheca resinae) and wall paint (Aureobasidium pullulans), and play their biggest role in recycling dead plant material. As biotrophs, they may form symbioses with algae (lichens), plant roots (mycorrhizae) or the leaves and stems of plants (endophytes). Other Ascomycota (Ceratocystis and Ophiostoma) form symbiotic associations with an array of arthropods, where they can line beetle galleries and provide nutrition for the developing larvae. In return, the beetles maintain a pure culture of the fungus and transport it to newly established galleries. As parasites, ascomycetes account for most of the animal and plant pathogens including Pneumocystis carinii, responsible for pneumonia of humans with compromised immune systems and Ophiostoma ulmi, the Dutch elm disease fungus that is responsible for the demise of elm trees in North America and Europe (Agrios, 1988).
Biogeography
Ascomycota can be found on all continents and many genera and species display a cosmopolitan distribution (Candida albicans or Aspergillus flavus). Others are found on more than one continent (Ophiostoma ulmi, or Cryphonectria parasitica), but many are known from only one narrowly restricted location. For example, the White Piedmont Truffle (Tuber magnatum) is known from only one province of Northern Italy.
Reproduction
From a human perspective, the most unusual aspect of all fungi is that they have more than one reproductive option. The textbook Ascomycota can make spores sexually (ascospores or meiospores) and asexually (condia or mitospores). Following meiosis, the ascospores take shape inside the ascus when new cell walls surround each nucleus as can be seen in the electron micrograph above (Wu and Kimbrough, 1992). Conidia contain mitotic nuclei, and their cell wall is simply a modified hyphal or yeast wall.

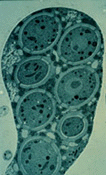
Ascus of a hyphal ascomycete (Pezizomycotina) as viewed by the electron microscope, © R. Vilgalys 1996.
Ascospores may or may not be shot by turgor pressure from the ascus and although wind is the primary dispersal agent once the spores have been released from the ascus, Ascomycota also use splashing or running water or animals to disperse their spores (Ingold, 1965). Conidial diversity reaches its climax with the Ascomycota, with forms ranging from single spores hardly different from hyphae(Geotrichum candidum) to elaborate heads of ornamented condida (Aspergillus niger) and beyond (Cole and Kendrick, 1981).
Life Cycle
Ascomycota are either single-celled (yeasts) or filamentous (hyphal) or both (dimorphic). Yeasts grow by budding or fission and hyphae grow apically and branch laterally. Most yeasts and filamentous Ascomycota are haploid, but some species, Saccharomyces cerevisiae for example, can also be diploid. Mitospores may simply reproduce the parent, or may also act as gametes to fertilize a compatible partner. Some Ascomycota must outbreed (heterothallic), others can also self, and some can only self (homothallic) (Alexopoulos et al. 1996).

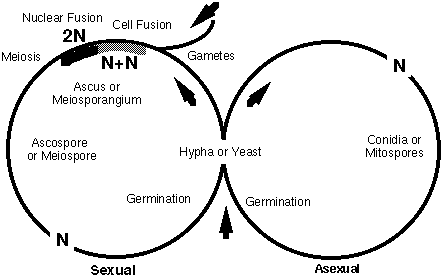
Life cycle of Ascomycota, © J. Taylor 1996
Genetic regulation of sex expression and mating is well-understood in some model Ascomycota such as yeast, where there are two sexes and mating is coordinated by oligopeptide pheromones (Marsh, 1991; Glass and Lorimer, 1991). In hyphal species, cytoplasmic fusion may not be immediately followed by nuclear fusion, leading to a short dikaryotic phase. The dikaryotic hyphae may be protected and nourished by differentiated haploid hyphae which form a fruiting body (the ascoma; plural ascomata). Ascomata may be closed (cleistothecium), open by a narrow orifice (perithecium), or broadly open like a cup (apothecium). Ascospores are released from the ascoma and germinate to form a new haploid mycelium.
Relationships of Ascomycota to other Fungi
The Ascomycota is sister group to the Basidiomycota. This relationship is supported by the presence in members of both phyla of cross-walls (septa) that divide the hypahe into segments, and pairs of unfused nuclei in these segments after mating and before nuclear fusion (dikaryons). Further support comes from the apparent homology between structures that coordinate simultaneous mitosis of the two dikaryotic nucli (Ascomycota croziers and Basidiomycota clamp-connections).
Discussion of Phylogenetic Relationships
Molecular phylogenetic analyses of nuclear and mitochondrial ribosomal RNA genes and protein coding genes support a monophyletic Ascomycota (Lutzoni et al 2004, James et al 2006, Spatafora et al 2006). Early diverging lineages of Ascomycota have been classified in Taphrinomycotina (Eriksson 2004; =Archiascomycetes Nishida and Sugiyama 1994). Due to the lack of strong support for the monophly of Taphrinomycotina (Nishida and Sugiyama 1994) and the paraphyletic resolution of these taxa in some analyses (Lutzoni et al 2004), Taphrinamycotina is not recognized in some classifications (Ericksson 2005). More recent analyses, however, that included multiple protein coding genes and RNA genes recovered a monophyletic Taphrinomycotina with greater support (James et al 2006, Liu et al 2006, Spatafora et al 2006).
The Taphrinomycotina includes yeast species (Pneumocystis, Schizosaccharomyces), dimorphic taxa (Taphrina spp.) and a filamentous sporocarp producing genus (Neolecta). The placement of Neolecta among the basal lineages of the Ascomycota is surprising because of the presence of an ascoma, a feature not found in the other basal lineages or in any Saccharomycotina (Landvik et al. 1992). However, there is no reason that the Saccharomycotina could not have lost ascomata as hyphal growth became suppressed in favor of yeasts. The Saccharomycotina form a well-supported monophyletic taxon, as do the Pezizomycotina (Gargas et al. 1995, Lutzoni et al 2004, Spatafora et al 2006). Asexual fungi sharing morphological or molecular characters of sexual Ascomycota are classified in the Ascomycota and its subtaxa; examples include Candida albicans (Saccharomycotina, Saccharomycetes) and Pencillium chrysogenum (Pezizomycotina, Eurotiomycetes).
By comparing nucleic acid sequences from 50 genes, the timing of Ascomycota evolution has been estimated, although results produced a wide geological time span depending on calibrations points used (Taylor and Berbee 2006). The Taphrinomycotina, Saccharomycotina and Pezizomycotina were likely established in the early Devonian, a bit more than 400 million years ago (mya). Some estimates, however, suggest a much earlier Ascomycota origin of ca. 1000 mya (Hedges et al 2001, Taylor and Berbee 2006). Fossils of early Ascomycota are not easy to recognize and the utility of some of them as exemplars of extant lineages is problematical (e.g., Paleopyrenomycites devonicus as a fossil Sordariomycetes). Thus, we still rely on generally accepted fossil dates external to Fungi (e.g., dicot-monocot split) for potentially more robust calibration points.
Subgroups of Ascomycota
The basal lineage or lineages of Ascomycota comprise four classes, Neolectomycetes, Pneumocystidomycetes, Schizosaccharomycetes, and Taphrinomycetes, which are classified in the subphylum, Taphrinomycotina (=Archiascomycetes Nishida and Sugiyama). The monophyly of Taphrinomycotina is questionable (Nishida and Sugiyama 1994, Eriksson 2005), but increased sampling of both taxa and genes has resulted in increased support for the monophyly of the taxon (James et al 2006, Liu et al 2006, Spatafora et al 2006).
The Saccharomycotina comprises the 'true yeasts' and is home to the most famous fungus, Saccharomyces cerevisiae, better known as the baker's yeast. Although most members are primarily unicellular, the basal taxa make abundant hyphae. The Saccharomycotina lack ascomata (Barnett et al., 1990). Current classification of the Saccharomycotina includes one class, Saccharomycetes, and one order, Saccharomycetales.
Pezizomycotina contain well over 90% of Ascomycota, and the species are hyphal, with almost all of the sexually reproducing forms possessing ascomata. The Pezizomycotina includes 11 well supported clades that are recognized as classes. Most of the recent molecular phylogenetic effort has been directed at this subphylum (reviewed in Lutzoni et al 2004, Spatafora et al 2006).
References
Agrios, G. N. 1988. Plant Pathology, third edition. Academic Press, San Diego.
Alexopoulos, C. J., C. W. Mims, and M. Blackwell. 1996. Introductory Mycology. John Wiley and Sons, New York. 868p.
Barnett, J. A., R. W. Payne, and D. Yarrow. 1990. Yeasts: characteristics and identification. Cambridge University Press, Cambridge.
Berbee, M. L., and J. W. Taylor. 1992a. Convergence in ascospore discharge mechanism among Pyrenomycete fungi based on 18S ribosomal RNA gene sequence. Mol. Phylog. Evol. 1:59-71.
Berbee, M. L., and J. W. Taylor. 1992b. Two ascomycete classes based on fruiting-body characters and ribosomal DNA sequence. Mol. Biol. Evol. 9:278-284.
Berbee, M. L., and J. W. Taylor. 1993. Dating the evolutionary radiations of the true fungi. Can. J. Bot. 71:1114-1127.
Bruns, T. D., R. Vilgalys, S. M. Barns, D. Gonzalez, D. S. Hibbett, D. J. Lane, L. Simon, S. Stickel, T. M. Szaro, W. G. Weisburg, and M. L. Sogin. 1992. Evolutionary relationships within the fungi: analyses of nuclear small subunit rRNA sequences. Mol. Phylog. Evol. 1:231-241.
Carroll, G.C. and D. T. Wicklow, 1992. The Fungal Community: Its Organization and Role in the Ecosystem. Marcel Dekker, Inc., New York.
Cole, G. T., and B. Kendrick. 1981. Biology of conidial fungi. Academic Press, New York.
Gargas, A., P. T. DePriest, M. Grube, and A. Tehler. 1995. Multiple origins of lichen symbioses in fungi suggested by SSU rDNA phylogeny. Science 268:1492-1495.
Glass, N. L., and I. A. J. Lorimer. 1991. Ascomycete mating types. Pages 193-216. in More gene manipulations in fungi (J. W. Bennett and L. L. Lasure, eds.). Academic Press, Orlando.
Griffin, D. H. 1994. Fungal Physiology. 2nd. Wiley-Liss, New York.
Heckman DS, Geiser DM, Eidell BR, Stauffer RL, Kardos NL, Hedges SB. 2001. Molecular evidence for the early colonization of land by fungi and plants. Science 293:1129-1133.
Ingold, C. T. 1965. Spore Liberation. Clarendon Press, Oxford.
James, T.Y., Kauff, F., Schoch, C., Matheny, P.B., Hofstetter, V., Cox, C.J., Celio, G., Gueidan, C., Fraker, E., Miadlikowska, J., Lumbsch, H.T., Rauhut, A., Reeb, V., Arnold, A.E., Amtoft, A., Stajich, J.E., Hosaka, K., Sung, G.-H., Johnson, D., ORourke, B., Crockett, M., Binder, M., Curtis, J.M., Slot, J.C., Wang, Z., Wilson, A.W., Schußler, A., Longcore, J.E., ODonnell, K., Mozley-Standridge, S., Porter, D., Letcher, P.M., Powell, M.J., Taylor, J.W., White, M.M., Griffith, G.W., Davies, D.R., Humber, R.A., Morton, J.B., Sugiyama, J., Rossman, A.Y., Rogers, J.D., Pfister, D.H., Hewitt, D., Hansen, K., Hambleton, S., Shoemaker, R.A., Kohlmeyer, J., Volkmann-Kohlmeyer, B., Spotts, R.A., Serdani, M., Crous, P.W., Hughes, K.W., Matsuura, K., Langer, E., Langer, G., Untereiner, W.A., Lucking, R., Budel, B., Geiser, D.M., Aptroot, A., Diederich, P., Schmitt, I., Schultz, M., Yahr, R., Hibbett, D.S., Lutzoni, F., McLaughlin, D.J., Spatafora, J.W. and Vilgalys, R. 2006. Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature (in press).
Landvik, S., O. E. Eriksson, A. Gargas, and P. Gustafsson. 1993. Relationships of the genus Neolecta (Neolectales ordo nov., Ascomycotina) inferred from 18s rDNA sequences. Syst. Ascomycetum 11:107-118.
Landvik, S., O. E. Eriksson, A. Gargas, and P. Gustafsson. 1993. Relationships of the genus Neolecta (Neolectales ordo nov., Ascomycotina) inferred from 18s rDNA sequences. Syst. Ascomycetum 11:107-118.
Liu, Y. J., M. C. Hodson and B. D. Hall. 2006. Loss of the flagellum happened only once in the fungal lineage: phylogenetic structure of Kingdom Fungi inferred from RNA polymerase II subunit genes. BMC Evolutionary Biology 2006, 6:74.
Lutzoni, F., F. Kauff, C. J. Cox, D. McLaughlin, G. Celio, B. Dentinger, M. Padamsee, D. Hibbett, T. Y. James, E. Baloch, M. Grube, V. Reeb, V. Hofstetter, C. Schoch, A. E. Arnold, J. Miadlikowska, J. Spatafora, D. Johnson, S. Hambleton, M. Crockett, R. Shoemaker, G.-H. Sung, R. Lücking, T. Lumbsch, K. O?Donnell, M. Binder, P. Diederich, D. Ertz, C. Gueidan, K. Hansen, R. C. Harris, K. Hosaka, Y.-W. Lim, B. Matheny, H. Nishida, D. Pfister, J. Rogers, A. Rossman, I. Schmitt, H. Sipman, J. Stone, J. Sugiyama, R. Yahr, and R. Vilgalys. 2004. Assembling the fungal tree of life: progress, classification, and evolution of subcellular traits. American Journal of Botany 91: 1446-1480.
Marsh, L. 1991. Signal transduction during pheromone response in yeast. Annu. Rev. Cell biol. 7:699-728.
Nishida, H., and J. Sugiyama. 1994. Archiascomycetes: Detection of a major new linage within the Ascomycota. Mycoscience 35:361-366.
Raju, N. B. 1992. Genetic control of the sexual cycle in Neurospora. Mycol. Res. 96:241-262,.
Spatafora, J. W. 1995a. Ascomal evolution among filamentous ascomycetes: evidence from molecular data. Can. J. Bot. S811-S815.
Spatafora, J., and M. Blackwell. 1993. Molecular systematics of unitunicate perithecial Ascomycetes. The Clavicipitales - Hypocreales connection. Mycologia 85:912-922.
Spatafora, J.W., Johnson, D., Sung, G.-H., Hosaka, K., ORourke, B., Serdani, M., Spotts, R., Lutzoni, F., Hofstetter, V., Fraker, E., Gueidan, C., Miadlikowska, J., Reeb, V., Lumbsch, T., Lücking, R., Schmitt, I., Aptroot, A., Roux, C., Miller, A., Geiser, D., Hafellner, J., Hestmark, G., Arnold, A.E., Büdel, B., Rauhut, A., Hewitt, D., Untereiner, W., Cole, M.S., Scheidegger, C., Schultz, M., Sipman., H. and Schoch, C. 2006. A five-gene phylogenetic analysis of the Pezizomycotina. Mycologia (in press).
Taylor, J. W. 1995. Making the Deuteromycota redundant: a practical integration of mitosporic and meiosporic fungi. Can. J. Bot. 73 (suppl.):s754-s759.
Taylor, J. W., B. Bowman, M. L. Berbee, and T. J. White. 1993. Fungal model organisms: phylogenetics of Saccharomyces, Aspergillus and Neurospora. Syst. Biol. 42:440-457.
Taylor, J. W. and M. L. Berbee. 2006. Dating divergences in the Fungal Tree of Life: Review and new analyses. Mycologia (in press).
Wessels, J. G. H. 1994. Developmental regulation of fungal cell wall formation. Ann. Rev. Phytopathol. 32:413-437.
Wu, C. G., and J. W. Kimbrough. 1992. Ultrastructural studies of ascosporogenesis in Ascobolus immersus. Mycologia 84:459-466.
edit internet links
Information on the Internet
- MYCONET. An international mycological journal mainly intended for the development of natural classifications of fungi and for the publication of related information
- Outline of Ascomycota - 2001. All accepted genera and taxa above the generic level in phylum Ascomycota.
- Freshwater Ascomycete Database. Carol Shearer, University of Illinois at Urbana-Champaign.
- LIAS. A Global Information System for Lichenized and Non-Lichenized Ascomycetes.
- Saccharomyces Genome Database.
- PezWeb. A Web site dedicated to the study of the Pezizales.
- Identifying Morels and False Morels.
- truffle.org. To promote research on truffle and ectomycorrhizae. At present the main emphasis of this project is to provide methods for the identification of truffles both at the morphological and molecular level. Istituto di Scienze Biochimiche, University of Parma.
- Truffles of the Great Basin. Robert Fogel, University of Michigan and Jack States, Northern Arizona University.
- The Truffle, Black Pearl of Perigord.
- Systematic Studies in Discomycetes: Pezizales. PEET Project. Harvard University.
- DERMBASE: Names of Dermateaceae (Ascomycetes). Botanischer Garten und Botanisches Museum Berlin-Dahlem, Freie Universität Berlin.
- The Aspergillus Website.
- Studies in the Lasiosphaeriaceae. NSF-PEET Project. The Field Museum, Chicago.
- Neurospora Genome Project. University of New Mexico.
- The Podospora anserina Web Page.
- Hypomyces. Descriptions, images, keys. Part of Monographic Studies of Hypocreales: Hypocrea and Hypomyces PEET Project. United States Department of Agriculture, Agricultural Research Service.
- Home of the Xylariaceae. Washington State University.
Title Illustrations

| Scientific Name | Taphrina |
|---|---|
| Comments | Asci of the peach leaf curl fungus atop a peach leaf |
| Body Part | asci |
| Copyright | © 1996 K. Wells |
| Scientific Name | Saccharomyces |
|---|---|
| Comments | Budding cells of the baker's and brewer's yeast |
| Copyright | © 1996 K. Wells |
| Scientific Name | Morchella |
|---|---|
| Comments | Two fruiting bodies (ascomata) of the edible morel with one sliced open |
| Body Part | fruiting bodies (ascomata) |
| Copyright |
© 1996 John W. Taylor

|
About This Page
Many thanks to Dave Carmean, Soren Rosendahl for scanning photos and David Maddison for page design advice.
John W. Taylor

University of California, Berkeley, California, USA
Joey Spatafora

Oregon State University, Corvallis, Oregon, USA
Mary Berbee

University of British Columbia, Vancouver, British Columbia, Canada
Correspondence regarding this page should be directed to John W. Taylor at
Page copyright © 2012 John W. Taylor, Joey Spatafora, and Mary Berbee
All Rights Reserved.
- First online 11 March 1996
- Content changed 09 October 2006
Citing this page:
Taylor, John W., Joey Spatafora, and Mary Berbee. 2006. Ascomycota. Sac Fungi. Version 09 October 2006 (under construction). http://tolweb.org/Ascomycota/20521/2006.10.09 in The Tree of Life Web Project, http://tolweb.org/













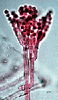
 Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site
Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site